Research
Department Antebi
Over the last several decades, studies in model genetic organisms have identified multiple evolutionarily conserved signalling pathways that regulate these processes, including insulin/IGF and mTOR signalling, mitochondrial function, dietary restriction mediated longevity, and signals from the reproductive system. Our lab has been instrumental in elucidating some of the key players at the nexus of these pathways, and ongoing projects include the dissection of nutrient sensing pathways and the role of metabolic signalling and reproduction in regulating animal lifespan.

Much of the work in the field has focused on pathway-specific mediators, but whether the different ageing-related pathways converge on common regulators or shared downstream processes largely remains an open question. Much of our recent work, therefore, has focused on deciphering convergent mechanisms to understand what lies at the heart of longevity and to identify clinical targets that could extend health- and lifespan. This area of research focuses on four main topics:
The nucleolus as a convergence point of longevity regulation
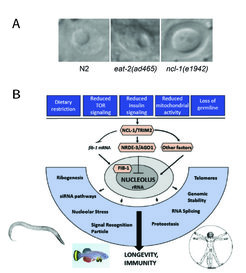
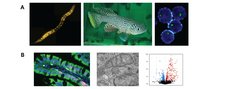
We have identified the nucleolus as a central convergence point of longevity regulation and found a number of genes involved in the regulation of nucleolar size and function. We have discovered that small nucleoli are a cellular hallmark of longevity not only in C. elegans but also in flies, fish, mice, and perhaps humans. The mechanisms by which the nucleolus contributes to ageing, however, remain unclear. Thus, unravelling whether and how different nucleolar functions (e.g. rRNA production and biogenesis, assembly of different ribonucleoprotein particles, splicing factors, siRNA pathways) affect longevity is a central research theme in our lab.
Dissecting how the MYC-MONDO and TFEB transcription factor network affects life span
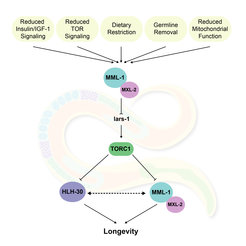
We have discovered an extensive helix-loop-helix (HLH) transcription factor network, consisting of the MML-1/MYC-MONDO complex and HLH-30/TFEB, which is required for life extension across multiple pathways. Among other things, this network largely regulates the metabolism of lipids, carbohydrates, amino acids, amines, nucleosides and mitochondria. How these processes actually relate to life span, however, remains to be elucidated. Our current work therefore aims to identify both upstream and downstream factors that regulate the MYC-MONDO and TFEB network.
Organellar communication and its role in ageing
Our current work highlights that different organelles (mitochondria, the endoplasmic reticulum, nucleoli and lysosomes) are not only involved in ageing, but that the crosstalk between them also contributes to immunity and longevity. Identifying the factors that mediate this communication and understanding how organelles cooperate to regulate lifespan is a major focus of our ongoing research.
Identification of metabolites that regulate animal health and life span
We have had a long-standing interest in understanding how naturally occurring metabolites can serve as signalling molecules that regulate animal health and life span, and are currently using metabolomic approaches to identify metabolites that impact longevity.
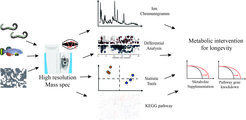
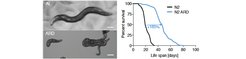
In addition, we are actively investigating different states of long-lived quiescence (called diapause) and how they relate to longevity. We study both dauer diapause, which occurs during the C.elegans third larval stage, as well as adult reproductive diapause (ARD), which animals enter in response to late larval food deprivation. Our current work focuses on unraveling the molecular and physiological pathways governing dauer diapause and ARD.





